Notre laboratoire fait partie du Genoscope et il est rattaché à
l'Institut de Biologie François Jacob du CEA. Le Commissariat à l'Enérgie Atomique et aux énérgies Alternatives est un acteur majeur de la recherche, du développement et de l'innovation dans quatre domaines principaux: défense et sécurité, énérgies peu carbonnées (nucléaire et énérgies renouvelables), recherche technologique pour l'industrie, recherche fondamentale en sciences physiques et sciences de la vie. Actuellement, le CEA dispose de plus de 20.000 employé(e)s réparti(e)s dans neuf centres situés en France.
Le Genoscope a été fondé en 1996 pour participer au projet de séquençage du génome Humain et développer des projets de génomiques en France avant de se tourner vers la génétique environnementale. Il fait partie de la Division de la Recherche Fondamentale (DRF, Fabrique de savoirs) depuis 2006 (Domaine Biologique). Le Genoscope dévelope des méthodes et des projets dédiés à l'exploitation de la biodiversité, notamment à l'aide de séquençage massif d'ADN et de bioinformatique. Il est ouvert à la communauté scientifique nationale à travers des appels à projets coordonnés dans le cadre de France Genomique depuis 2012. Les projets couvrent toute la biodiversité, plus particulièrement la génomique des plantes, des champignons ou du plancton mais aussi la métagénomique d'écosystèmes complexes. Le Genoscope est rattaché à l'Université de Paris-Saclay qui est l'une des meilleures universités française en Europe, placée 13ème dans la classement de Shanghai 2021 et reconnue pour la qualité de ses programmes éducatifs mais aussi de son équipe enseignante. L'université est fière de sa visibilité internationale, forte de la réputation de ses 275 laboratoires de recherche et de leurs équipes qui proposent un exceptionnel soutien quotidien pour l'intégration et le développement de 65.000 étudiants multiculturels.
Le Genoscope est composé de plusieurs laboratoires dotés à la fois d'équipements de séquençage et de support informatique. Le laboratoire de séquençage maîtrise les technologies de courtes et longues lectures (Illumina, MGI et Oxford Nanopore) et a la capacité de gérer des projets génomiques de grande ampleur avec de très nombreux échantillons. Depuis 2012, le Genoscope gère et séquence les échantillons des expéditions TARA Ocean. Le Genoscope dispose d'un centre de calculs de 1700 cœurs avec plusieurs nœuds forts d'une mémoire de 2 à 6 To ainsi qu'un espace de stockage distribué d'1,5 Po. De plus, le Genoscope a accès au centre de calculs du CEA. ( Le CCRT doté d'une infrastructure de calcul à grande échelle et d'un stockage de 5 Po )
Our lab is part of the Genoscope and attached to the Institut de Biologie François Jacob of the CEA. The French Alternative Energies and Atomic Energy Commission is a key player in research, development and innovation in four main areas: defense and security, low carbon energies (nuclear and renewable energies), technological research for industry, fundamental research in the physical sciences and life sciences. Currently the CEA has more than 20,000 employees and is established in nine centers spread throughout France.
Genoscope was founded in 1996 to contribute to the Human genome project and develop genomic programs in France and has subsequently turned toward environmental genomics. It has been part of the CEA Fundamental Research Division (DRF, The Knowledge Factory) since 2006 (Biology field). Genoscope is developing methods and projects for the exploitation of biodiversity, in particular with respect to massive DNA sequencing and bioinformatics. It has been open to the national scientific community through calls for coordinated projects in the context of France Genomique since 2012. The projects cover all biodiversity, particularly the genomics of plants, fungi, plankton and the metagenomics of complex ecosystems. Genoscope is affiliated to the Paris-Saclay university which is one of the leading French and European universities, rated 13th in the 2021 Shanghai ranking and recognised for the quality of both its educational programmes and teaching staff. The university also boasts high international visibility thanks to the reputation of its 275 research laboratories and their teams and provides outstanding daily support for the integration and development of 65,000 multicultural students.
Genoscope is composed of several research laboratories with both sequencing and IT equipment. The sequencing laboratory operates short- and long-reads technologies (Illumina, MGI and Oxford Nanopore) and has the capacity to operate large-scale genomic projects with a high number of samples. Since 2012, Genoscope has managed and sequenced the samples from the TARA Ocean expeditions. Genoscope incorporates a 1700-core computing cluster with several large-memory nodes (2-6Tb), and a globally distributed storage of 1.5 PB. In addition, the Genoscope has access to the CEA computing infrastructure. ( CCRT with dedicated large-scale computing infrastructure and storage of 5 PB )
The main activities of the LBGB is to develop and evaluate new bioinformatics technologies and software to be used in original and large-scale genomic projects, and in particular with the goals of generating chromosome-scale assemblies of complex genomes based on a combination of long-reads sequencing with long-range information; of providing a gene annotation platform for eukaryotic genomes; and of performing comparative genomics analyzes aiming to establish links between the specificities of a given genome and its life traits. Here are several topics we are currently working on:
L'activité principale du LBGB est d'évaluer les nouvelles technologies et de développer des logiciels de bioinformatique utilisés dans des projets originaux d'envergure et plus particulièrement dans le but de produire des assemblages à l'échelle des chromosomes de génomes complexes basés sur l'association de séquençage longues lectures et d'informations 'long-range'; de proposer une plateforme d'annotation de gènes pour les génomes eucaryotes; et de produire des analyses de génomique comparative dans le but d'établir des liens entre les particularités d'un génome et les conditions de vie de l'organisme étudié. Voici quelques sujets sur lesquels nous travaillons actuellement:
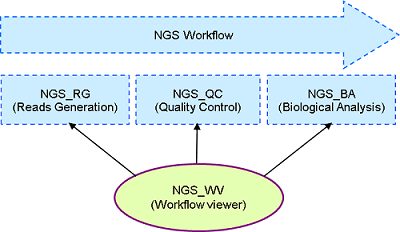
En relation avec le laboratoire de séquençage, nous évaluons en permanence les technologies de séquençage et les protocoles associés. Cette veille technologique nous permet de proposer des stratégies de séquençage adaptées aux projets du Genoscope.
In connection with the sequencing lab, we continuously evaluate the sequencing technologies and their associated protocols. This technological survey allows us to propose sequencing strategies adapted to Genoscope projects. We are also developing bioinformatic tools to check the quality of sequencing data produced at Genoscope.
Les standards évoluent rapidement et les assemblages à l'échelle des chromosomes tout comme les annotations nécessitent des méthodes à la pointe des technologies. Les premiers échantillons du projet Tara Oceans ont généré une grande proportion de séquences inconnues soulignant la nécessité de générer des banques de référence plus complète de génomes d'organismes marins. Nous développons de nouvelles méthodes dans le but d'obtenir des génomes complets d'organismes inconnus, un objectif encore inaccessible pour les eucaryotes.
Standards are evolving rapidly, and chromosome-level assemblies, as well as annotations integrating state-of-the-art methods are needed. The first sequencing of the Tara Oceans project generated a high proportion of unknown sequences, showing the strong need to generate a more complete database of marine organisms. We are developing new methodologies with the final goal of obtaining near-complete genomes of unknown organisms, presently an unreached goal for eukaryotes.
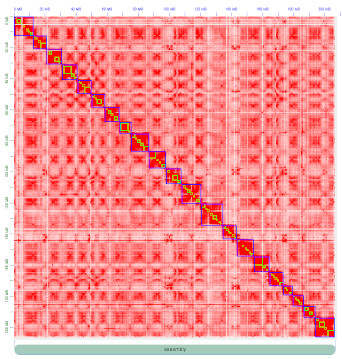
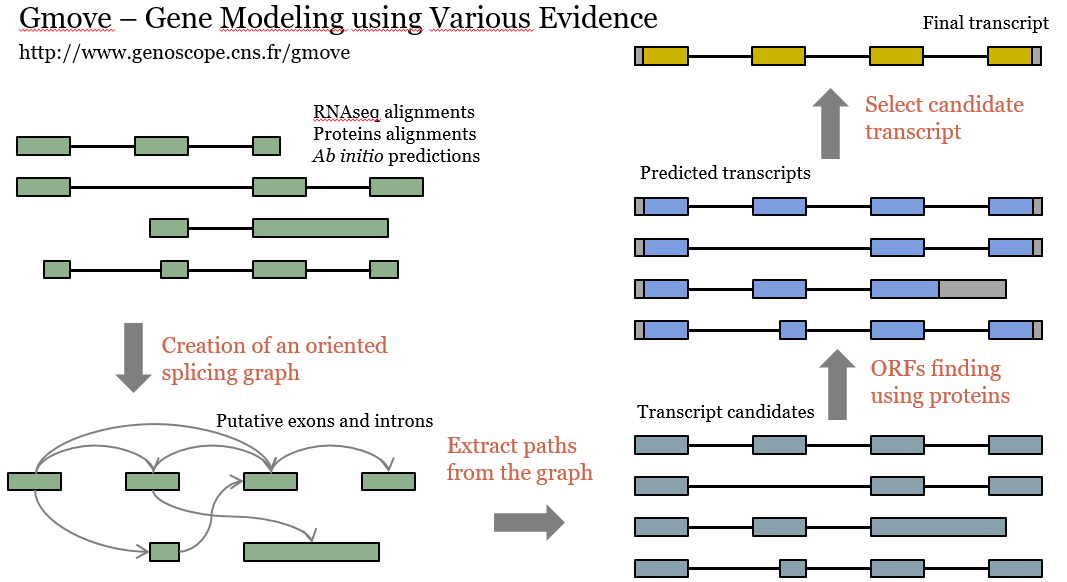
De plus avec la diminution des coûts de séquençage, une grande variété de génomes sera séquencée dans les années à venir, avec l'objectif de produire un assemblage de référence pour chaque espèce. Le goulot d'étranglement sera alors la prédiction de gènes, c'est pourquoi nous travaillons au développement d'un prédicteur de gène appelé «Gmove», capable de faire de la prédiction de novo mais aussi de transférer des annotations d'un génome à un autre.
Moreover, with the dropoff of the sequencing costs, we could expect that a large variety of genomes will be resequenced, with the goal of generating several references assemblies for a given species. One bottleneck will be the gene prediction, and for this we are working on the development of a gene predictor, called «Gmove», that can be used to perform de novo gene prediction as well as to transfer annotations from one genome to another.
Il a pour but de séquencer le génome de 4500 espèces marines eucaryotes comprenant : mollusques, crustacés, annélides, cnidaires, ascidies, algues unicellulaires et pluricellulaires, éponges et poissons, soit environ un tiers des espèces marines connues de l’hexagone et des territoires ultramarins. Les données récoltées seront déposées dans des bases de données en libre accès pour la communauté scientifique et compléteront ainsi les inventaires de biodiversité.
Its aim is to sequence the genomes of 4,500 eukaryotic marine species (species whose cells have a nucleus), including molluscs, crustaceans, annelids, cnidarians, ascidians, unicellular and multicellular algae, sponges and fish, i.e. around a third of known marine species in France and the overseas territories. The data collected will be deposited in an open-access database for the scientific community, thus completing biodiversity inventories.
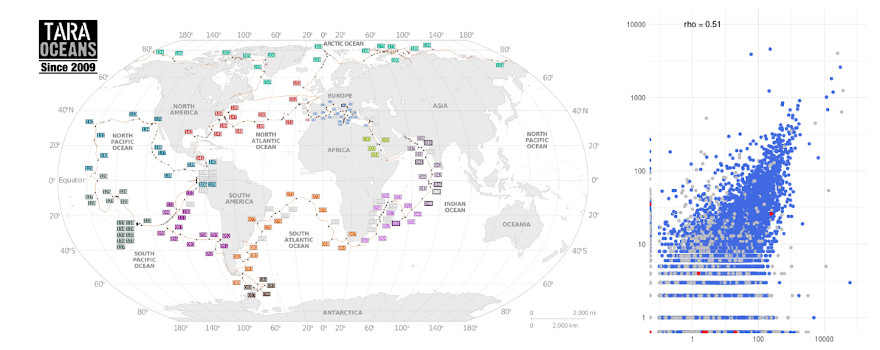
Développements spécifiques dans le domaine de la transcriptomique pour intégrer les possibilités du séquençage nanopore de molécules d’ARN. Nous prévoyons de créer des outils de construction de cartes de transcrits avec leurs profils d'expression au sein d'une expérience. Ces développements seront adaptés aux données environmentales produites par les divers consortia avec qui nous collaborons.
Specific developments in the field of transcriptomics to integrate the new possibilities of nanopore sequencing using RNA molecules. We plan to create tools for building complete transcript maps and their associated expression profiles across experiments, adapted to the environmental datasets produced by the different consortia with whom we are collaborating.
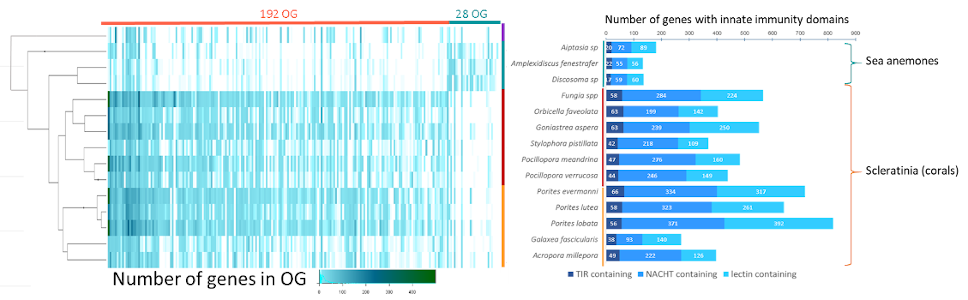
Les organismes sessiles à longue durée de vie doivent persister face à un large éventail de menaces biotiques et abiotiques tout au long de leur vie. Nous avons étudié les caractéristiques génomiques associées à une telle longévité en séquençant, assemblant et annotant les génomes de plusieurs espèces. Nous avons ensuite utilisé le nombre croissant de séquences génomiques complètes pour étudier l'évolution convergente des caractéristiques génomiques potentiellement en lien avec la longévité.
Long-lived sessile organisms must persist in the face of a wide range of abiotic and biotic threats over their lifespans. We investigated the genomic features associated with such a long lifespan by sequencing, assembling and annotating genomes of several species. We then used the growing number of whole-genome sequences to investigate the parallel evolution of genomic characteristics potentially underpinning longevity.
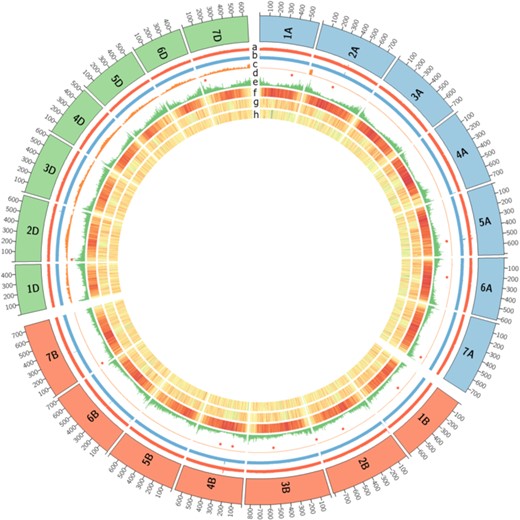
L'analyse génomique comparative nécessite des outils de visualisation. Pour cela, les assemblages et les annotations génomiques sont accessibles via une interface dédiée basée sur le Generic Genome Browser. Nous développons également des outils spécifiques qui nous permettent d'étudier la synténie entre les génomes et leur histoire évolutive.
Comparative genomic analysis requires visualization tools, for that purpose, assemblies and genomic features are available through a dedicated interface based on the Generic Genome Browser. We also develop specific tools that allow us to investigate synteny between genomes and evolutionary history of studied genomes.
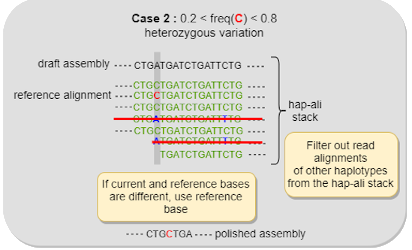
Le développement des outils est guidé par les applications scientifiques du Genoscope. Nous utilisons généralement des logiciels existants développés par d'autres groupes de bioinformatique, mais nous devons gérer les problématiques bioinformatiques liées à nos propres projets scientifiques. Les outils disponibles ne sont pas forcément adaptés à nos besoins ; nous devons les évaluer, les modifier et, éventuellement, en développer de nouveaux.
Tools development is guided by the scientific applications of Genoscope. Generally, we use existing software developed by other bioinformatic groups, but we have to manage the bioinformatic issues brought by our own scientific projects. Available tools are not necessarily adapted to our needs, we have to evaluate these tools, modify them and eventually develop new tools.

Responsable d'équipe Team leader

Assemblage de génome Genome assembly

Assemblage de génome Genome assembly

Analyse transcriptomique Transcriptomic analysis

Génomique comparative Comparative genomics

Assemblage de génome Genome assembly

Génétique comparative Comparative genomics

Production de données de séquençage Sequencing data production

Thèse en metatranscriptomique Thesis on metatranscriptomics long reads within the LongTREC network

Production de données de séquençage Sequencing data production

Assemblage de génome Genome assembly

Assemblage de génome Genome assembly

Production de données de séquençage Sequencing data production

Production de données de séquençage Sequencing data production

Prédiction de gènes Gene prediction

Génomique comparative Comparative genomics

Assemblage de génome Genome assembly

Prédiction de g&eagrave;nes Gene prediction

Prédiction de g&eagrave;nes Gene prediction
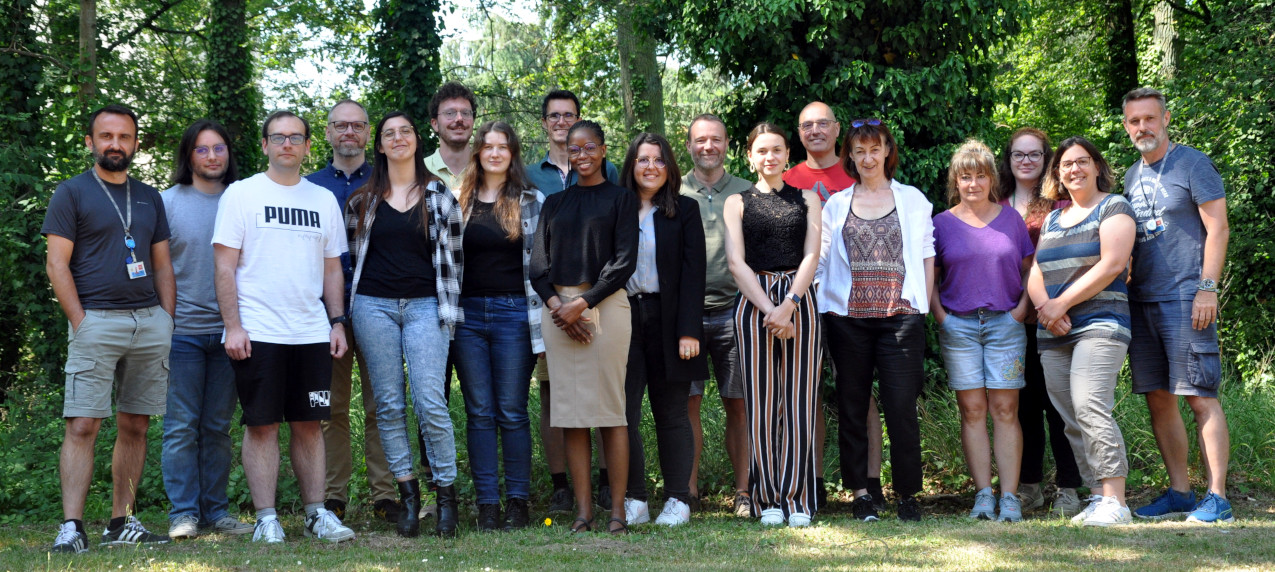

Chaque année, nous accueillons plusieurs étudiants pour effectuer leur stage dans notre laboratoire. Si vous êtes intéressé(e) par un stage dans l'un de nos domaines scientifiques, n'hésitez pas à postuler en nous envoyant votre CV à: stage_lbgb@genoscope.cns.fr
Each year, we welcome several students to do their internship in our laboratory. If you are interested in doing your internship in one of our scientific fields, do not hesitate to apply by sending us a CV to: stage_lbgb@genoscope.cns.fr
jmaury@genoscope.cns.fr
01 60 87 25 00
